Medical image datasets
TorchIO offers tools to easily download publicly available datasets from different institutions and modalities.
The interface is similar to torchvision.datasets.
If you use any of them, please visit the corresponding website (linked in each description) and make sure you comply with any data usage agreement and you acknowledge the corresponding authors' publications.
If you would like to add a dataset here, please open a discussion on the GitHub repository.
CT-RATE
CtRate
CtRate
Bases: SubjectsDataset
CT-RATE dataset.
This class helps loading the CT-RATE dataset , which contains chest CT scans with associated radiology reports and abnormality labels.
The dataset must have been downloaded previously.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
root
|
TypePath
|
Root directory where the dataset has been downloaded. |
required |
split
|
TypeSplit
|
Dataset split to use, either |
'train'
|
num_subjects
|
int | None
|
Optional limit on the number of subjects to load (useful for
debugging). If |
None
|
report_key
|
str
|
Key to use for storing radiology reports in the Subject metadata. |
'report'
|
sizes
|
list[int] | None
|
List of image sizes (in-plane, in voxels) to include. |
None
|
load_fixed
|
bool
|
If |
True
|
verify_paths
|
bool
|
If |
False
|
**kwargs
|
Additional arguments for SubjectsDataset. |
{}
|
Examples:
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
IXI
IXI
IXI
Bases: SubjectsDataset
Full IXI dataset.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
root
|
TypePath
|
Root directory to which the dataset will be downloaded. |
required |
transform
|
An instance of
|
required | |
download
|
bool
|
If set to |
False
|
modalities
|
Sequence[str]
|
List of modalities to be downloaded. They must be in
|
('T1', 'T2')
|
Warning
The size of this dataset is multiple GB.
If you set download to True, it will take some time
to be downloaded if it is not already present.
Examples:
>>> import torchio as tio
>>> transforms = [
... tio.ToCanonical(), # to RAS
... tio.Resample((1, 1, 1)), # to 1 mm iso
... ]
>>> ixi_dataset = tio.datasets.IXI(
... 'path/to/ixi_root/',
... modalities=('T1', 'T2'),
... transform=tio.Compose(transforms),
... download=True,
... )
>>> print('Number of subjects in dataset:', len(ixi_dataset)) # 577
>>> sample_subject = ixi_dataset[0]
>>> print('Keys in subject:', tuple(sample_subject.keys())) # ('T1', 'T2')
>>> print('Shape of T1 data:', sample_subject['T1'].shape) # [1, 180, 268, 268]
>>> print('Shape of T2 data:', sample_subject['T2'].shape) # [1, 241, 257, 188]
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
IXITiny
IXITiny
Bases: SubjectsDataset
This is the dataset used in the main notebook. It is a tiny version of IXI, containing 566 \(T_1\)-weighted brain MR images and their corresponding brain segmentations, all with size \(83 \times 44 \times 55\).
It can be used as a medical image MNIST.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
root
|
TypePath
|
Root directory to which the dataset will be downloaded. |
required |
transform
|
Transform | None
|
An instance of
|
None
|
download
|
bool
|
If set to |
False
|
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
EPISURG
EPISURG
EPISURG
Bases: SubjectsDataset
EPISURG is a clinical dataset of \(T_1\)-weighted MRI from 430 epileptic patients who underwent resective brain surgery at the National Hospital of Neurology and Neurosurgery (Queen Square, London, United Kingdom) between 1990 and 2018.
The dataset comprises 430 postoperative MRI. The corresponding preoperative MRI is present for 268 subjects.
Three human raters segmented the resection cavity on partially overlapping subsets of EPISURG.
If you use this dataset for your research, you agree with the Data use agreement presented at the EPISURG entry on the UCL Research Data Repository and you must cite the corresponding publications.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
root
|
TypePath
|
Root directory to which the dataset will be downloaded. |
required |
transform
|
Transform | None
|
An instance of
|
None
|
download
|
bool
|
If set to |
False
|
Warning
The size of this dataset is multiple GB.
If you set download to True, it will take some time
to be downloaded if it is not already present.
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
get_labeled()
Get dataset from subjects with manual annotations.
get_unlabeled()
Get dataset from subjects without manual annotations.
get_paired()
Get dataset from subjects with pre- and post-op MRI.
Kaggle datasets
RSNAMICCAI
RSNAMICCAI
Bases: SubjectsDataset
RSNA-MICCAI Brain Tumor Radiogenomic Classification challenge dataset.
This is a helper class for the dataset used in the
RSNA-MICCAI Brain Tumor Radiogenomic Classification challenge hosted on
kaggle . The dataset must be downloaded before
instantiating this class (as opposed to, e.g., torchio.datasets.IXI).
This kaggle kernel includes a usage example including preprocessing of all the scans.
If you reference or use the dataset in any form, include the following citation:
U.Baid, et al., "The RSNA-ASNR-MICCAI BraTS 2021 Benchmark on Brain Tumor Segmentation and Radiogenomic Classification", arXiv:2107.02314, 2021.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
root_dir
|
TypePath
|
Directory containing the dataset ( |
required |
train
|
bool
|
If |
True
|
ignore_empty
|
bool
|
If |
True
|
Examples:
>>> import torchio as tio
>>> from subprocess import call
>>> call('kaggle competitions download -c rsna-miccai-brain-tumor-radiogenomic-classification'.split())
>>> root_dir = 'rsna-miccai-brain-tumor-radiogenomic-classification'
>>> train_set = tio.datasets.RSNAMICCAI(root_dir, train=True)
>>> test_set = tio.datasets.RSNAMICCAI(root_dir, train=False)
>>> len(train_set), len(test_set)
(582, 87)
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
RSNACervicalSpineFracture
RSNACervicalSpineFracture
Bases: SubjectsDataset
RSNA 2022 Cervical Spine Fracture Detection dataset.
This is a helper class for the dataset used in the RSNA 2022 Cervical Spine Fracture Detection hosted on kaggle . The dataset must be downloaded before instantiating this class.
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
MNI
ICBM2009CNonlinearSymmetric
ICBM2009CNonlinearSymmetric
Bases: SubjectMNI
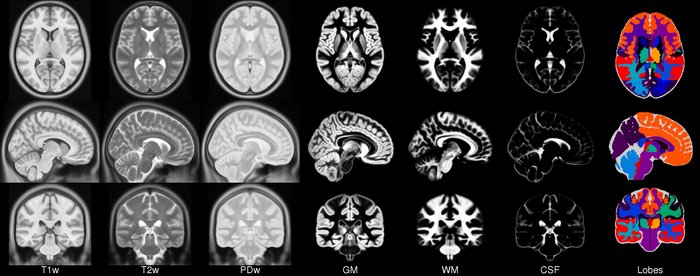
ICBM template.
More information can be found in the website .
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
load_4d_tissues
|
bool
|
If |
True
|
Examples:
>>> import torchio as tio
>>> icbm = tio.datasets.ICBM2009CNonlinearSymmetric()
>>> icbm
ICBM2009CNonlinearSymmetric(Keys: ('t1', 'eyes', 'face', 'brain', 't2', 'pd', 'tissues'); images: 7)
>>> icbm = tio.datasets.ICBM2009CNonlinearSymmetric(load_4d_tissues=False)
>>> icbm
ICBM2009CNonlinearSymmetric(Keys: ('t1', 'eyes', 'face', 'brain', 't2', 'pd', 'gm', 'wm', 'csf'); images: 9)
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
Colin27
Colin27
Bases: SubjectMNI
shape
property
spatial_shape
property
spacing
property
NAME_TO_LABEL = {name: label for label, name in (TISSUES_2008.items())}
class-attribute
instance-attribute
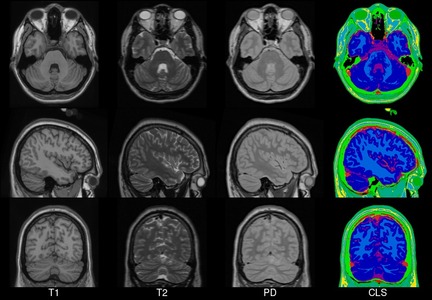
Colin27 MNI template.
More information can be found in the website of the 1998 and 2008 versions.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
version
|
Template year. It can be |
required |
Warning
The resolution of the 2008 version is quite high. The
subject instance will contain four images of size
\(362 \times 434 \times 362\), therefore applying a transform to
it might take longer than expected.
Examples:
>>> import torchio as tio
>>> colin_1998 = tio.datasets.Colin27(version=1998)
>>> colin_1998
Colin27(Keys: ('t1', 'head', 'brain'); images: 3)
>>> colin_1998.load()
>>> colin_1998.t1
ScalarImage(shape: (1, 181, 217, 181); spacing: (1.00, 1.00, 1.00); orientation: RAS+; memory: 27.1 MiB; type: intensity)
>>>
>>> colin_2008 = tio.datasets.Colin27(version=2008)
>>> colin_2008
Colin27(Keys: ('t1', 't2', 'pd', 'cls'); images: 4)
>>> colin_2008.load()
>>> colin_2008.t1
ScalarImage(shape: (1, 362, 434, 362); spacing: (0.50, 0.50, 0.50); orientation: RAS+; memory: 217.0 MiB; type: intensity)
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
Pediatric
Pediatric
Bases: SubjectMNI
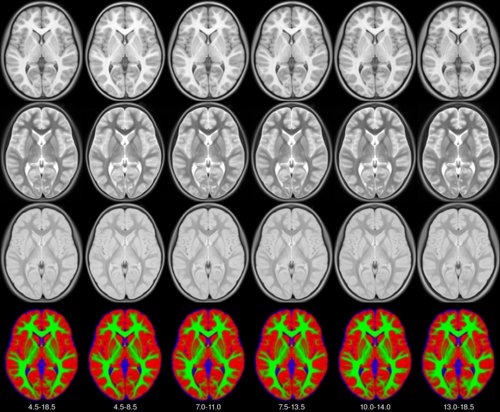
MNI pediatric atlases.
See the MNI website for more information.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
years
|
Tuple of 2 ages. Possible values are: |
required | |
symmetric
|
If |
False
|
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
Sheep
Sheep
Bases: SubjectMNI
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
BITE3
BITE3
Bases: BITE
Pre- and post-resection MR images in BITE.
The goal of BITE is to share in vivo medical images of patients wtith brain tumors to facilitate the development and validation of new image processing algorithms.
Please check the BITE website for more information and acknowledgments instructions.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
root
|
TypePath
|
Root directory to which the dataset will be downloaded. |
required |
transform
|
Transform | None
|
An instance of
|
None
|
download
|
bool
|
If set to |
False
|
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
ITK-SNAP
BrainTumor
BrainTumor
Bases: SubjectITKSNAP
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
T1T2
T1T2
Bases: SubjectITKSNAP
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
AorticValve
AorticValve
Bases: SubjectITKSNAP
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
3D Slicer
Slicer
Slicer
Bases: Subject
Sample data provided by 3D Slicer .
See the Slicer wiki for more information.
For information about licensing and permissions, check the Sample Data module .
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
name
|
One of the keys in |
'MRHead'
|
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
FPG
FPG
FPG
Bases: Subject
3T \(T_1\)-weighted brain MRI and corresponding parcellation.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
load_all
|
bool
|
If |
False
|
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.
MedMNIST
OrganMNIST3D
OrganMNIST3D
Bases: MedMNIST
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
NoduleMNIST3D
NoduleMNIST3D
Bases: MedMNIST
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
AdrenalMNIST3D
AdrenalMNIST3D
Bases: MedMNIST
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
FractureMNIST3D
FractureMNIST3D
Bases: MedMNIST
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
VesselMNIST3D
VesselMNIST3D
Bases: MedMNIST
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
SynapseMNIST3D
SynapseMNIST3D
Bases: MedMNIST
from_batch(batch)
classmethod
Instantiate a dataset from a batch generated by a data loader.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
batch
|
dict
|
Dictionary generated by a data loader, containing data that can be converted to instances of [Subject][.torchio.Subject]. |
required |
dry_iter()
set_transform(transform)
Set the transform attribute.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
transform
|
Callable | None
|
Callable object, typically an subclass of
|
required |
ZonePlate
ZonePlate
ZonePlate
Bases: Subject
Synthetic data generated from a zone plate.
The zone plate is a circular diffraction grating that produces concentric rings of light and dark bands. This dataset is useful for testing image processing algorithms, particularly those related to frequency analysis and interpolation.
See equation 10.63 in Practical Handbook on Image Processing for Scientific Applications by Bernd Jähne.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
size
|
int
|
The size of the generated image along all dimensions. |
501
|
shape
property
spatial_shape
property
spacing
property
get_inverse_transform(warn=True, ignore_intensity=False, image_interpolation=None)
Get a reversed list of the inverses of the applied transforms.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
warn
|
bool
|
Issue a warning if some transforms are not invertible. |
True
|
ignore_intensity
|
bool
|
If |
False
|
image_interpolation
|
str | None
|
Modify interpolation for scalar images inside transforms that perform resampling. |
None
|
apply_inverse_transform(**kwargs)
Apply the inverse of all applied transforms, in reverse order.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
**kwargs
|
Keyword arguments passed on to
|
{}
|
check_consistent_attribute(attribute, relative_tolerance=1e-06, absolute_tolerance=1e-06, message=None)
Check for consistency of an attribute across all images.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
attribute
|
str
|
Name of the image attribute to check |
required |
relative_tolerance
|
float
|
Relative tolerance for |
1e-06
|
absolute_tolerance
|
float
|
Absolute tolerance for |
1e-06
|
Examples:
>>> import numpy as np
>>> import torch
>>> import torchio as tio
>>> scalars = torch.randn(1, 512, 512, 100)
>>> mask = torch.tensor(scalars > 0).type(torch.int16)
>>> af1 = np.eye([0.8, 0.8, 2.50000000000001, 1])
>>> af2 = np.eye([0.8, 0.8, 2.49999999999999, 1]) # small difference here (e.g. due to different reader)
>>> subject = tio.Subject(
... image = tio.ScalarImage(tensor=scalars, affine=af1),
... mask = tio.LabelMap(tensor=mask, affine=af2)
... )
>>> subject.check_consistent_attribute('spacing') # no error as tolerances are > 0
Note
To check that all values for a specific attribute are close
between all images in the subject, numpy.allclose() is used.
This function returns True if
\(|a_i - b_i| \leq t_{abs} + t_{rel} * |b_i|\), where
\(a_i\) and \(b_i\) are the \(i\)-th element of the same
attribute of two images being compared,
\(t_{abs}\) is the absolute_tolerance and
\(t_{rel}\) is the relative_tolerance.
get_image(image_name)
Get a single image by its name.
load()
Load images in subject on RAM.
unload()
Unload images in subject.
add_image(image, image_name)
Add an image to the subject instance.
remove_image(image_name)
Remove an image from the subject instance.