RandomLabelsToImage
RandomLabelsToImage
Bases: RandomTransform, IntensityTransform
Randomly generate an image from a segmentation.
Based on the work by Billot et al.: A Learning Strategy for Contrast-agnostic MRI Segmentation and Partial Volume Segmentation of Brain MRI Scans of any Resolution and Contrast.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
label_key
|
str | None
|
String designating the label map in the subject that will be used to generate the new image. |
None
|
used_labels
|
Sequence[int] | None
|
Sequence of integers designating the labels used
to generate the new image. If categorical encoding is used,
|
None
|
image_key
|
str
|
String designating the key to which the new volume will be saved. If this key corresponds to an already existing volume, missing voxels will be filled with the corresponding values in the original volume. |
'image_from_labels'
|
mean
|
Sequence[TypeRangeFloat] | None
|
Sequence of means for each label.
For each value \(v\), if a tuple \((a, b)\) is
provided then \(v \sim \mathcal{U}(a, b)\).
If |
None
|
std
|
Sequence[TypeRangeFloat] | None
|
Sequence of standard deviations for each label.
For each value \(v\), if a tuple \((a, b)\) is
provided then \(v \sim \mathcal{U}(a, b)\).
If |
None
|
default_mean
|
TypeRangeFloat
|
Default mean range. |
(0.1, 0.9)
|
default_std
|
TypeRangeFloat
|
Default standard deviation range. |
(0.01, 0.1)
|
discretize
|
bool
|
If |
False
|
ignore_background
|
bool
|
If |
False
|
**kwargs
|
See |
{}
|
Tip
It is recommended to blur the new images in order to simulate
partial volume effects at the borders of the synthetic structures. See
RandomBlur.
Examples:
>>> import torchio as tio
>>> subject = tio.datasets.ICBM2009CNonlinearSymmetric()
>>> # Using the default parameters
>>> transform = tio.RandomLabelsToImage(label_key='tissues')
>>> # Using custom mean and std
>>> transform = tio.RandomLabelsToImage(
... label_key='tissues', mean=[0.33, 0.66, 1.], std=[0, 0, 0]
... )
>>> # Discretizing the partial volume maps and blurring the result
>>> simulation_transform = tio.RandomLabelsToImage(
... label_key='tissues', mean=[0.33, 0.66, 1.], std=[0, 0, 0], discretize=True
... )
>>> blurring_transform = tio.RandomBlur(std=0.3)
>>> transform = tio.Compose([simulation_transform, blurring_transform])
>>> transformed = transform(subject) # subject has a new key 'image_from_labels' with the simulated image
>>> # Filling holes of the simulated image with the original T1 image
>>> rescale_transform = tio.RescaleIntensity(
... out_min_max=(0, 1), percentiles=(1, 99)) # Rescale intensity before filling holes
>>> simulation_transform = tio.RandomLabelsToImage(
... label_key='tissues',
... image_key='t1',
... used_labels=[0, 1]
... )
>>> transform = tio.Compose([rescale_transform, simulation_transform])
>>> transformed = transform(subject) # subject's key 't1' has been replaced with the simulated image
See also
__call__(data)
Transform data and return a result of the same type.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
data
|
InputType
|
Instance of |
required |
get_base_args()
Provides easy access to the arguments used to instantiate the base class
(Transform) of any transform.
This method is particularly useful when a new transform can be represented as a variant
of an existing transform (e.g. all random transforms), allowing for seamless instantiation
of the existing transform with the same arguments as the new transform during apply_transform.
Note
The p argument (probability of applying the transform) is excluded to avoid
multiplying the probability of both existing and new transform.
add_base_args(arguments, overwrite_on_existing=False)
Add the init args to existing arguments
validate_keys_sequence(keys, name)
staticmethod
Ensure that the input is not a string but a sequence of strings.
to_hydra_config()
Return a dictionary representation of the transform for Hydra instantiation.
arguments_are_dict()
Check if main arguments are dict.
Return True if the type of all attributes specified in the
args_names have dict type.
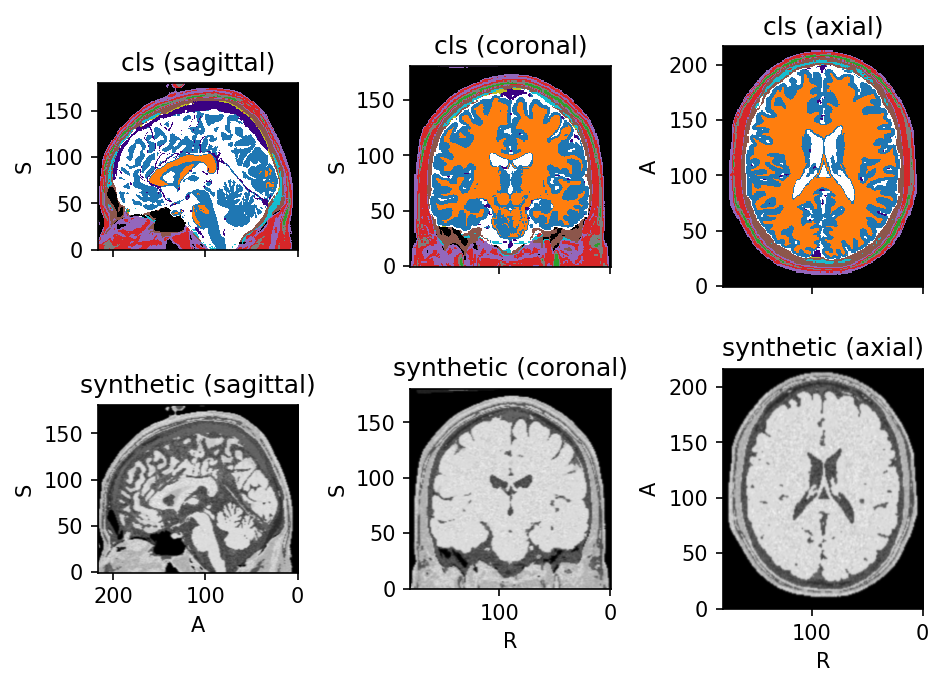
Source code
import torch
import torchio as tio
torch.manual_seed(42)
colin = tio.datasets.Colin27(2008)
label_map = colin.cls
colin.remove_image('t1')
colin.remove_image('t2')
colin.remove_image('pd')
downsample = tio.Resample(1)
blurring_transform = tio.RandomBlur(std=0.6)
create_synthetic_image = tio.RandomLabelsToImage(
image_key='synthetic',
ignore_background=True,
)
transform = tio.Compose((
downsample,
create_synthetic_image,
blurring_transform,
))
colin_synth = transform(colin)
colin_synth.plot()