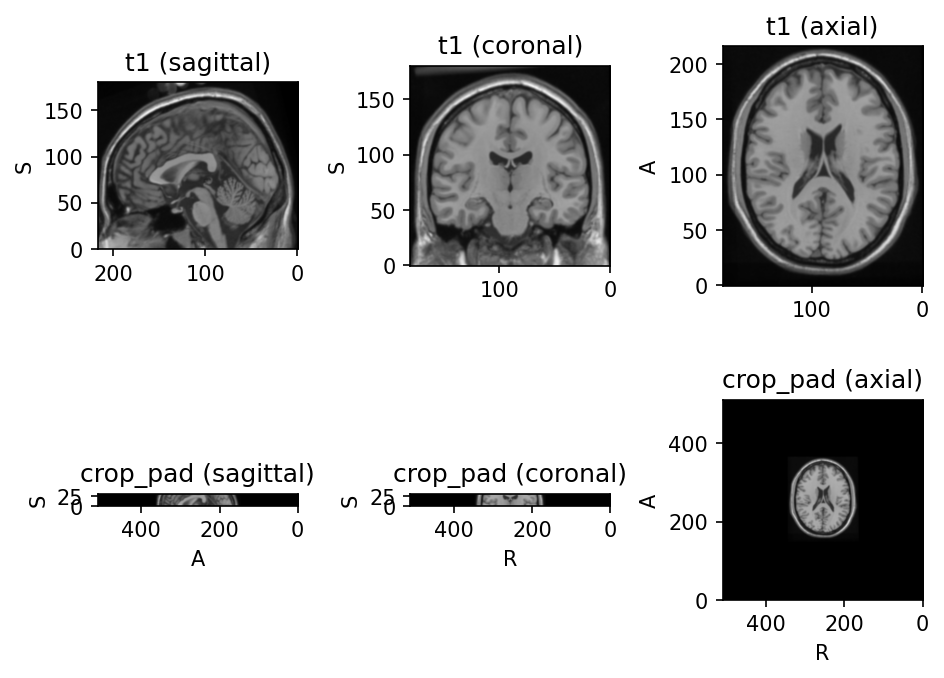
CropOrPad
CropOrPad
Bases: SpatialTransform
Modify the field of view by cropping or padding to match a target shape.
This transform modifies the affine matrix associated to the volume so that physical positions of the voxels are maintained.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
target_shape
|
int | TypeTripletInt | None
|
Tuple \((W, H, D)\). If a single value \(N\) is
provided, then \(W = H = D = N\). If |
None
|
padding_mode
|
str | float
|
Same as |
0
|
mask_name
|
str | None
|
If |
None
|
labels
|
Sequence[int] | None
|
If a label map is used to generate the mask, sequence of labels to consider. |
None
|
only_crop
|
bool
|
If |
False
|
only_pad
|
bool
|
If |
False
|
**kwargs
|
See |
{}
|
Examples:
>>> import torchio as tio
>>> subject = tio.Subject(
... chest_ct=tio.ScalarImage('subject_a_ct.nii.gz'),
... heart_mask=tio.LabelMap('subject_a_heart_seg.nii.gz'),
... )
>>> subject.chest_ct.shape
torch.Size([1, 512, 512, 289])
>>> transform = tio.CropOrPad(
... (120, 80, 180),
... mask_name='heart_mask',
... )
>>> transformed = transform(subject)
>>> transformed.chest_ct.shape
torch.Size([1, 120, 80, 180])
Warning
If target_shape is None, subjects in the dataset
will probably have different shapes. This is probably fine if you are
using patch-based training .
If you are using full volumes for training and a batch size larger than
one, an error will be raised by the DataLoader
while trying to collate the batches.
_get_six_bounds_parameters(parameters)
staticmethod
Compute bounds parameters for ITK filters.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
parameters
|
ndarray
|
Tuple \((w, h, d)\) with the number of voxels to be cropped or padded. |
required |
Returns:
| Type | Description |
|---|---|
TypeSixBounds
|
Tuple \((w_{ini}, w_{fin}, h_{ini}, h_{fin}, d_{ini}, d_{fin})\), |
TypeSixBounds
|
where \(n_{ini} = \left \lceil \frac{n}{2} \right \rceil\) and |
TypeSixBounds
|
\(n_{fin} = \left \lfloor \frac{n}{2} \right \rfloor\). |
Examples: