RemapLabels
RemapLabels
Bases: LabelTransform
Modify labels in a label map.
Masking can be used to split the label into two during the inverse transformation .
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
remapping
|
dict[int, int]
|
Dictionary that specifies how labels should be remapped. The keys are the old labels, and the corresponding values replace them. |
required |
masking_method
|
TypeMaskingMethod
|
Defines a mask for where the label remapping is applied. It can be one of:
|
None
|
**kwargs
|
See |
{}
|
Examples:
>>> import torch
>>> import torchio as tio
>>> def get_image(*labels):
... tensor = torch.as_tensor(labels).reshape(1, 1, 1, -1)
... image = tio.LabelMap(tensor=tensor)
... return image
...
>>> image = get_image(0, 1, 2, 3, 4)
>>> remapping = {1: 2, 2: 1, 3: 1, 4: 7}
>>> transform = tio.RemapLabels(remapping)
>>> transform(image).data
tensor([[[[0, 2, 1, 1, 7]]]])
Warning
The transform will not be correctly inverted if one of the values in
remapping is also in the input image:
>>> tensor = torch.as_tensor([0, 1]).reshape(1, 1, 1, -1)
>>> subject = tio.Subject(label=tio.LabelMap(tensor=tensor))
>>> mapping = {3: 1} # the value 1 is in the input image
>>> transform = tio.RemapLabels(mapping)
>>> transformed = transform(subject)
>>> back = transformed.apply_inverse_transform()
>>> original_label_set = set(subject.label.data.unique().tolist())
>>> back_label_set = set(back.label.data.unique().tolist())
>>> original_label_set
{0, 1}
>>> back_label_set
{0, 3}
Examples:
>>> import torchio as tio
>>> # Target label map has the following labels:
>>> # {
>>> # 'left_ventricle': 1, 'right_ventricle': 2,
>>> # 'left_caudate': 3, 'right_caudate': 4,
>>> # 'left_putamen': 5, 'right_putamen': 6,
>>> # 'left_thalamus': 7, 'right_thalamus': 8,
>>> # }
>>> transform = tio.RemapLabels({2:1, 4:3, 6:5, 8:7})
>>> # Merge right side labels with left side labels
>>> transformed = transform(subject)
>>> # Undesired behavior: The inverse transform will remap ALL left side labels to right side labels
>>> # so the label map only has right side labels.
>>> inverse_transformed = transformed.apply_inverse_transform()
>>> # Here's the *right* way to do it with masking:
>>> transform = tio.RemapLabels({2:1, 4:3, 6:5, 8:7}, masking_method="Right")
>>> # Remap the labels on the right side only (no difference yet).
>>> transformed = transform(subject)
>>> # Apply the inverse on the right side only. The labels are correctly split into left/right.
>>> inverse_transformed = transformed.apply_inverse_transform()
__call__(data)
Transform data and return a result of the same type.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
data
|
InputType
|
Instance of |
required |
get_base_args()
Provides easy access to the arguments used to instantiate the base class
(Transform) of any transform.
This method is particularly useful when a new transform can be represented as a variant
of an existing transform (e.g. all random transforms), allowing for seamless instantiation
of the existing transform with the same arguments as the new transform during apply_transform.
Note
The p argument (probability of applying the transform) is excluded to avoid
multiplying the probability of both existing and new transform.
add_base_args(arguments, overwrite_on_existing=False)
Add the init args to existing arguments
validate_keys_sequence(keys, name)
staticmethod
Ensure that the input is not a string but a sequence of strings.
to_hydra_config()
Return a dictionary representation of the transform for Hydra instantiation.
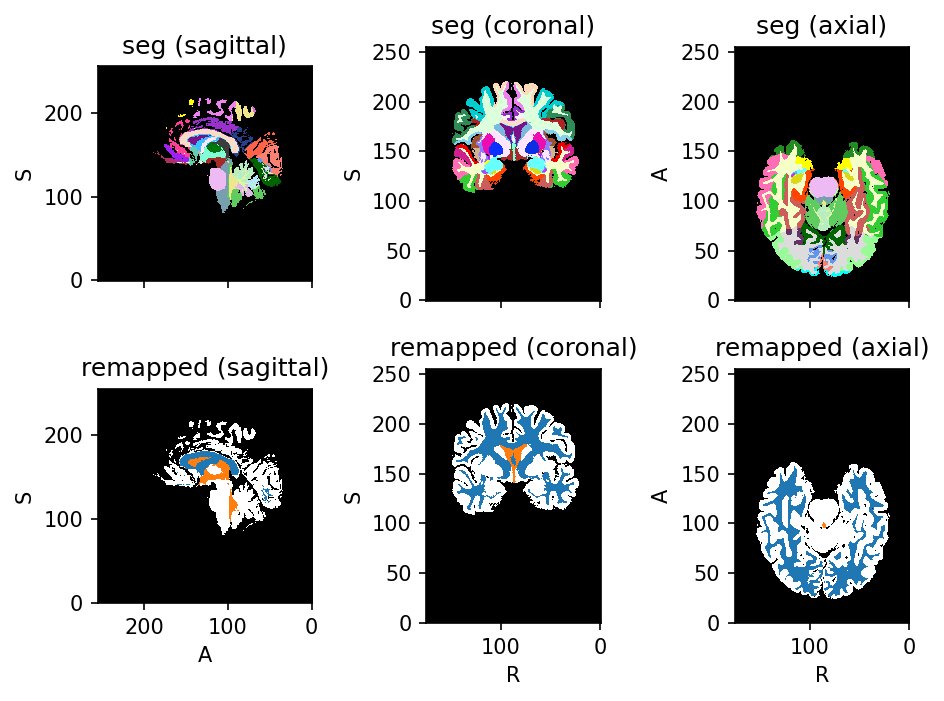
Source code
import torchio as tio
subject = tio.datasets.FPG()
subject.remove_image('t1')
background_labels = (0, 1, 2, 3, 4)
csf_labels = (5, 12, 16, 47, 52, 53)
white_matter_labels = (
45, 46,
66, 67,
81, 82,
83, 84,
85, 86,
87,
89, 90,
91, 92,
93, 94,
)
not_gray_matter_labels = (
background_labels
+ csf_labels
+ white_matter_labels
)
gray_matter_labels = [
label for label in subject.GIF_COLORS
if label not in not_gray_matter_labels
]
labels_groups = (
background_labels,
gray_matter_labels,
white_matter_labels,
csf_labels,
)
remapping = {}
for target, labels in enumerate(labels_groups):
for label in labels:
remapping[label] = target
parcellation_to_tissues = tio.RemapLabels(remapping)
tissues = parcellation_to_tissues(subject).seg
subject.add_image(tissues, 'remapped')
subject.plot()